Hacking Explainability: Using LIME as a General Framework for Complex Tasks
Deep learning has undeniably transformed medical image analysis. Yet, as architectures grow more sophisticated (e.g., see our recent blogpost about U-Mamba) , the ability to understand “why” models make specific decisions lags behind. The medical field strongly demands transparency. However, standard eXplainable Artificial Intelligence (XAI) often leaves us with more questions than answers.
The Limitation of Explainability Methods to Classification
One core issue is the strict limitation of explainability methods to classification tasks. Methods like Grad-CAM [1] or Integrated Gradients [2] answer basic questions perfectly. They easily answer: “The model says there is a tumor. Where?”
However, they usually produce blurry, imprecise heatmaps. When you need precision and answering complex questions like “What makes the model believe that a tumor is there? Is it just voxel intensity or the spatial context of the tumor?” these methods fall short.
So, how do we bridge this gap? We go back to the basics and “hack” one of the foundational methods in the XAI space: Local Interpretable Model-Agnostic Explanations (LIME).
The Intuition and History Behind LIME
When LIME was introduced in 2016 [3], it was a massive success and a pivotal early method in the XAI timeline. Before the ecosystem became saturated with highly complex mathematical frameworks, LIME offered something brilliant: simplicity.
The core intuition is exceptionally easy to understand. Thus, imagine a model that classifies animals. If we take an image of a dog and systematically hide its ears, then its tail, and then its legs, does the model still think it’s a dog? By observing how the prediction changes as we hide parts of the input, we learn what features the model relies on.
In a medical context, if an ML model segments a malignant cell, we can ask: does the prediction collapse if the high-density nucleus is obscured, or does it rely more on the texture of the surrounding cytoplasm?
While newer methods, including ones based on Shapley values [4], provide robust, axiomatic guarantees based on cooperative game theory, the complexity of underlying math makes it difficult to adapt to non-standard real-world problems. LIME’s simplicity is exactly what makes it so “hackable” for complex tasks like medical image segmentation.
How LIME Works – In Detail
To effectively hack LIME, we must understand its internal mechanics. Let’s look at a practical, tutorial-style example. We want our ML model to classify whether an image contains cell structures.
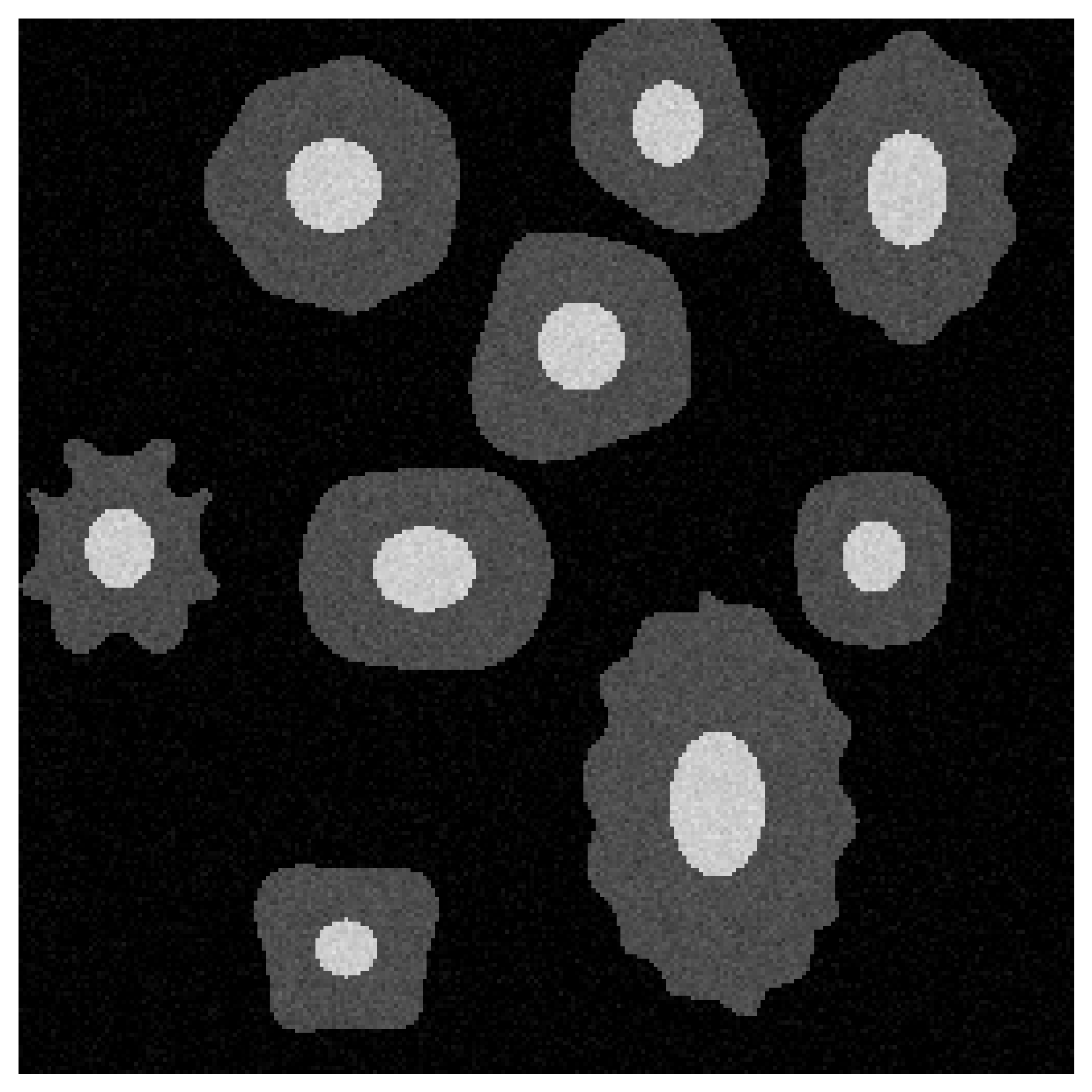
Figure 1: Raw cellular structures present in a medical image.
Figure 1 shows our raw, unedited input image. Our ML model’s task here is image classification. It must successfully identify if target cells are present. To explain this classification, we first define a local “neighborhood” around our image.
Intuitively, a neighbor is a of slightly altered version of the original image. Think of testing how someone recognizes a specific car. We could show them photos missing the front bumper, trunk, or headlights. In our medical example, the neighborhood consists of hundreds of these tweaked images.
To create these variations, we first group adjacent pixels into meaningful anatomical regions called superpixels. The Simple Linear Iterative Clustering (SLIC) [5] algorithm is typically used for this step. SLIC balances two main factors: spatial compactness and color proximity. If compactness is high, we get rigid, square blocks. Conversely, if color proximity dominates, boundaries become incredibly erratic.
Figure 2: Cells grouped into anatomically logical superpixels using SLIC.
Look closely at Figure 2 to see this balance. Notice the distinct rounding around the circular cell nuclei. The superpixels naturally follow the actual biological tissue borders.
Next, hundreds of perturbed image variations are generated. To create these, a random fraction of superpixels—typically ranging from 10% to 50%—is perturbed in each neighbor. For instance, values of perturbed superpixels might be replaced with random noise or color negation. Perturbation strategies can be tailored for the medical domain to ensure substitution plausibility. For example, instead of blacking out a region, we can swap non-enhanced regions for contrast-enhanced ones or even replace soft tissue with the intensity values of dense bone.
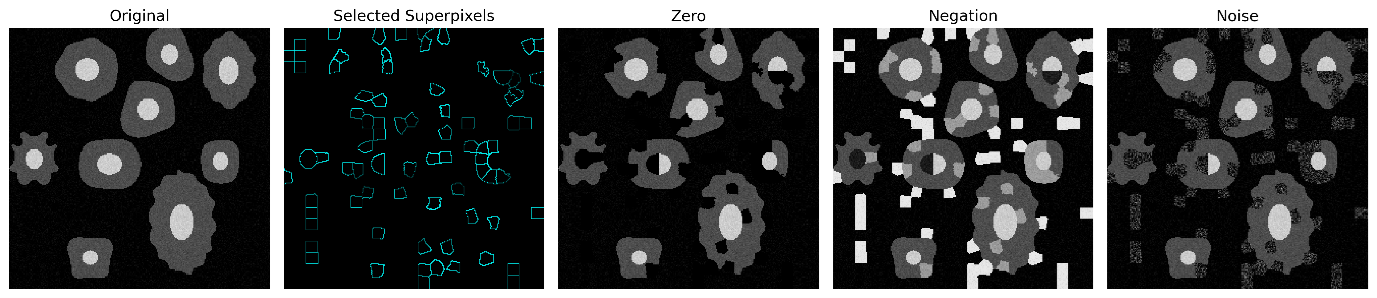
Figure 3: Different perturbation strategies: Zeroing, Negation, and Gaussian Noise.
Then, every perturbed image is passed through a complex AI model. Output probabilities are systematically recorded.
To analyze how these probabilities fluctuate based on the absence of specific superpixels, LIME creates a simple surrogate model, denoted as g. This surrogate model completely ignores raw pixels in the original image x. Instead, it only checks the presence or absence of specific superpixels, represented by π. It treats each superpixel like a simple on/off switch.
To train this surrogate, LIME minimizes the following loss:
Where h is the complex model, g is the interpretable surrogate, and π represents the ‘on/off’ vector of superpixels. Weights are assigned to each neighbor x’ based on its similarity to the original image x, ensuring the surrogate model g focuses on the most relevant local changes. By minimizing the loss L, the surrogate model g allows to deduce exactly which superpixels drove the ML model’s decision. The resulting coefficients from this simple model become our final feature importance scores.
Adapting LIME for Semantic Segmentation Tasks
Semantic segmentation assigns a specific class to every single pixel. The primary goal is strict pixel-level attribution. An example of such task is segmentation of nuclei, which is visualized for the analyzed instance in Figure 3. In such setup, explanations must reveal exactly what features drove these precise boundaries.
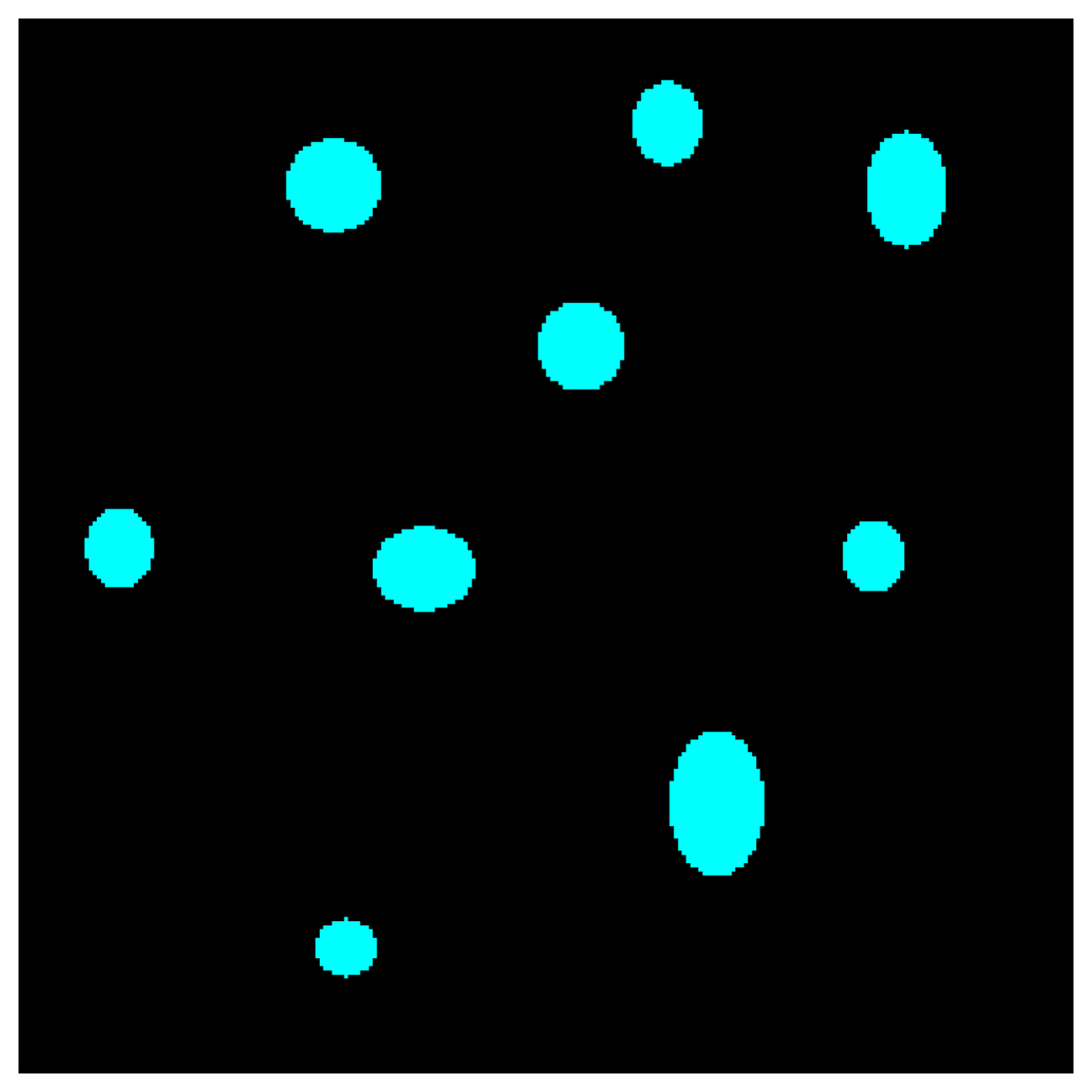

Figure 4: Semantic segmentation mask generated by ML model.
Standard LIME expects a single output probability for classification. However, adapting LIME for semantic segmentation requires accounting for segmentation mask, where each voxel is treated as an individual prediction.
To adapt the loss function, we redefine the target variable. Instead of a single class probability, we use a scalar value s representing the model’s retention of the target mask:
Where M is the set of pixels in the original predicted mask, and h(x′)i i is the model’s confidence for the target class at pixel i in the perturbed image. The surrogate model g is then trained to minimize:
This allows the surrogate model to quantify exactly how much of the original segmented area “survives” when specific superpixels are perturbed.
These explanations provide actionable insights by validating different model behaviors. Consider two AI models trained for the exact same task. Generated LIME maps clearly expose their distinct internal logic paths: Figure 5 shows that Model A relies solely on nuclei color and hence the model is not robust. Contrary, Figure 6 shows that Model B accounts for the spatial context of the cytoplasm, making it more robust to variations in color.
The approach described can be even further refined, to explain segmentation of a single nucleus to get even finer grained explanation. In such case, the target loss function should be modified to approximate changes in number of pixels classified only in the region where predicted in the beginning.
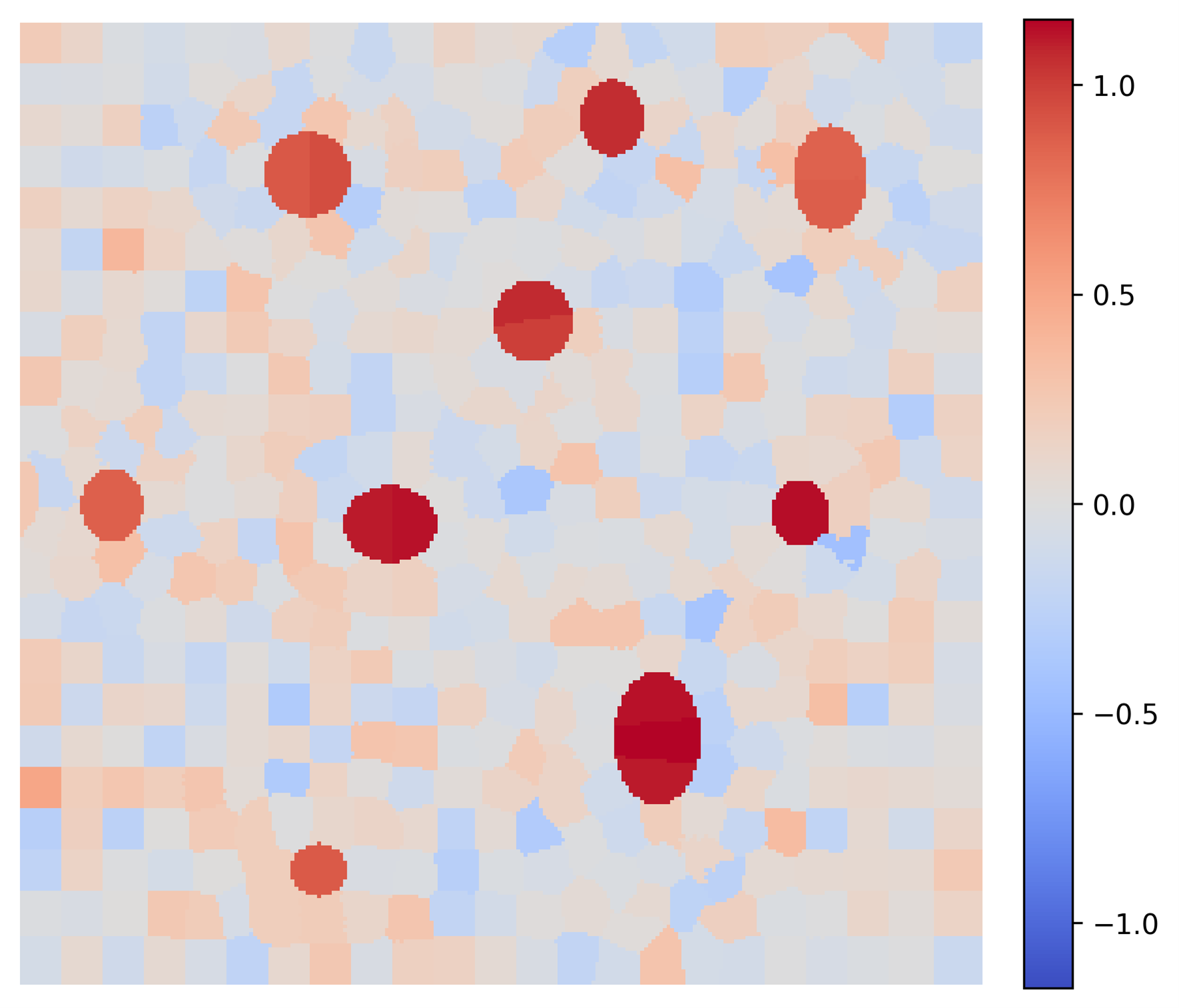
Figure 5: Explanation of Segmentation Given by Model A: Relying incorrectly solely on nuclei hue heuristics.
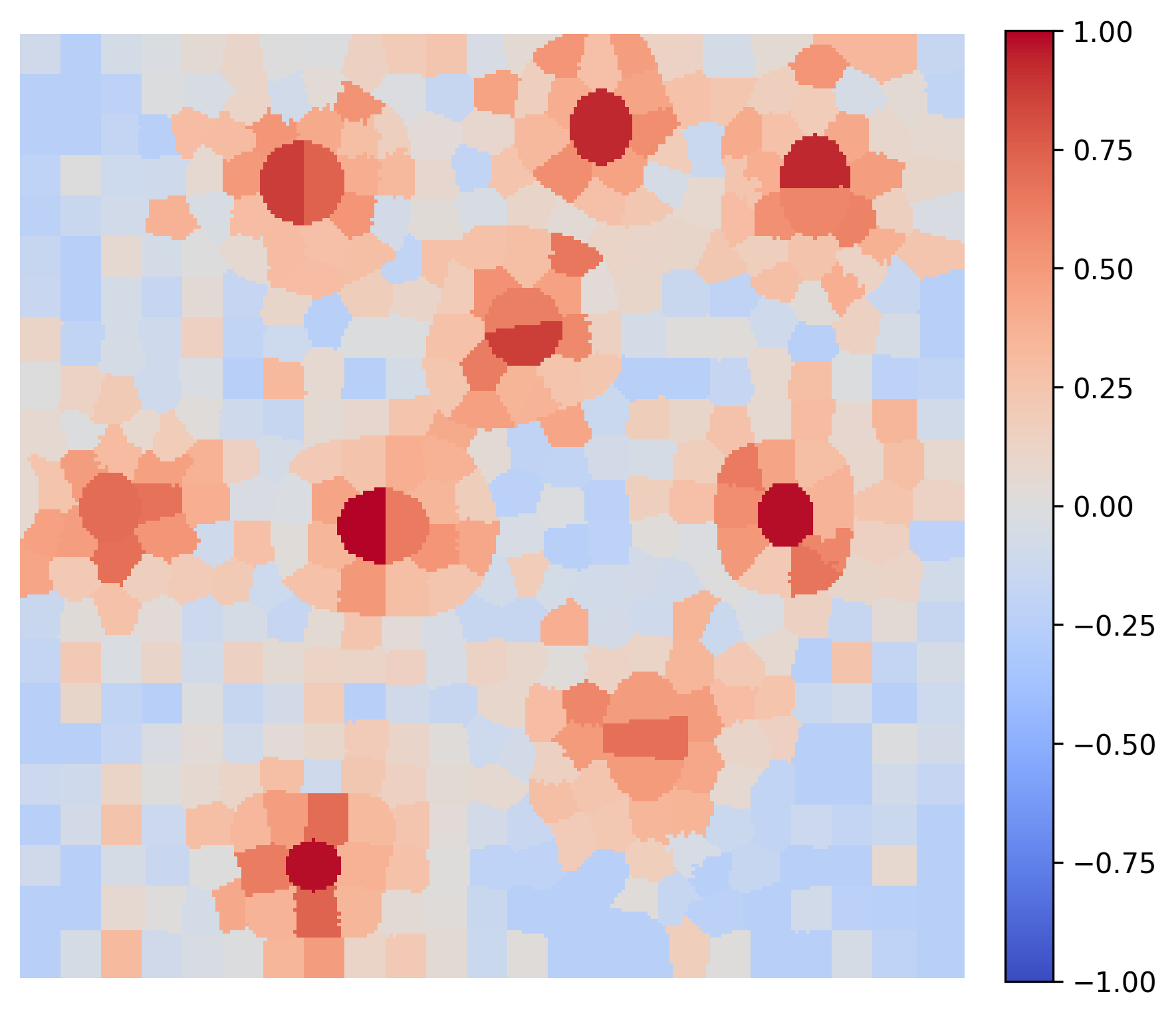
Figure 6: Explanation of Segmentation Given by Model B: Relying correctly on both nuclei hue and surrounding spatial context.
Conclusion: Purpose-Driven Explainability
A common trap in modern ML engineering is doing explainability just to “check a box”. When researchers apply XAI without a clear hypothesis, they often struggle to draw actionable conclusions from the resulting heatmaps.
Every attempt at explainability should be driven by a specific clinical or engineering need. Are you trying to prove the model isn’t looking at surgical artifacts? Are you trying to verify if it uses contextual tissue to classify a tumor?
LIME is not perfect. It is known to be sensitive to the chosen neighborhood size and can sometimes exhibit instability across multiple runs. Applying LIME with different perturbation strategies can lead to drastically different results. However, when you understand the mechanics beneath it, LIME’s degrees of freedom allow you to set up an experiment that can answer almost any targeted question. Moreover, used wisely, it is no longer just an out-of-the-box tool and becomes a highly adaptable framework for dissecting complex medical models.
Bibliography
- Selvaraju, R.R. et al. (2017) ‘Grad-CAM: Visual Explanations From Deep Networks via Gradient-Based Localization’, in Proceedings of the IEEE International Conference on Computer Vision (ICCV). https://doi.org/10.1007/s11263-019-01228-7
- Sundararajan, M., Taly, A. and Yan, Q. (2017) ‘Axiomatic attribution for deep networks’, in Proceedings of the 34th International Conference on Machine Learning – Volume 70 (ICML’17), pp. 3319–3328. https://dl.acm.org/doi/10.5555/3305890.3306024
- Ribeiro, M.T., Singh, S. and Guestrin, C. (2016) ‘“Why Should I Trust You?”: Explaining the Predictions of Any Classifier’, in Proceedings of the 22nd ACM SIGKDD International Conference on Knowledge Discovery and Data Mining, pp. 1135–1144. https://doi.org/10.1145/2939672.2939778
- Lundberg, S.M. and Lee, S.-I. (2017) ‘A unified approach to interpreting model predictions’, in Proceedings of the 31st International Conference on Neural Information Processing Systems (NIPS’17), pp. 4768–4777. https://dl.acm.org/doi/10.5555/3295222.3295230
- Achanta, R. et al. (2012) ‘SLIC Superpixels Compared to State-of-the-Art Superpixel Methods’, IEEE Trans. Pattern Anal. Mach. Intell., 34(11), pp. 2274–2282. https://doi.org/10.1109/TPAMI.2012.120